Software
Maptcha (download)
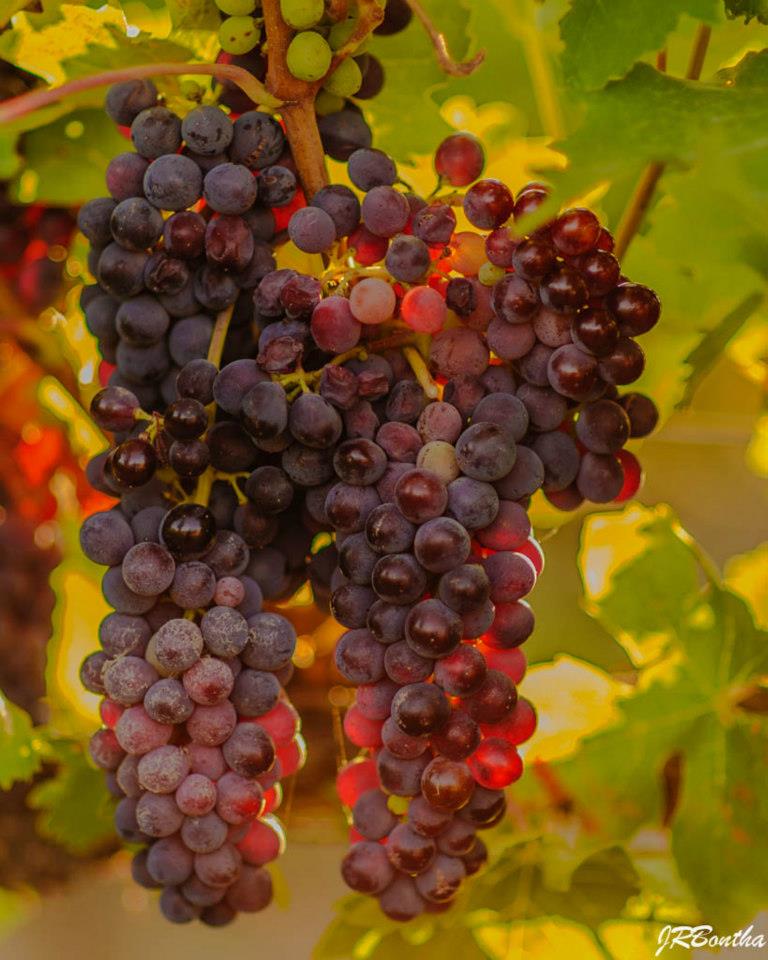
Parallel workflow for hybrid genome scaffolding (e.g., for combining contigs with long reads).
Citations:
Bhowmik, Oieswarya, Tazin Rahman, and Ananth Kalyanaraman. "Maptcha:
An Efficient Parallel Workflow for hybrid genome scaffolding." BMC
Bioinformatics, vol. 25, no. 263, 2024. DOI: 10.1186/s12859-024-05878-4.
JEM-Mapper (download)
Parallel sketch-based mapping tool for long read workflows.
Citations:
Rahman, Tazin, Oieswarya Bhowmik, and Ananth Kalyanaraman. "An
Efficient Parallel Sketch-based Algorithmic Workflow for Mapping Long Reads."
IEEE/ACM Transactions on Computational Biology and Bioinformatics, vol. 22, no. 1, pp. 13-26, 2025. DOI: 10.1109/TCBB.2024.3489478.
Contact network dashboard for DASON hospital data (download)
This codebase is used for generating the visual dashboard of mixing matrices, for patients across different units in the DASON database. The dashboard shows different aggregate measured corresponding to patient mixing from hospitals.
Citations:
K. Madhobi, A. Kalyanaraman, D.J. Anderson, E. D. Ashley, R. K.
Moehring, E. Lofgren. JAMA Network Open, doi:
10.1001/jamanetworkopen.2022.25508
Delta-screening dynamic community detection
(download)
Dynamic community detection using the ?-screening technique, integrated using the Louvain code for community detection. Implemented in C/C++.
Citations:
N. Zarayeneh, A. Kalyanaraman, IEEE TNSE, 2021.
N. Zarayeneh, A. Kalyanaraman, ASONAM'19, pp.9-16, 2019.
PaKman(download)
Parallel genome assembler for scalable generation of genomic contigs. Implemented in C/C++ and runs on distributed memory clusters (MPI).
Citations:
P. Ghosh, S. Krishnamoorthy, A. Kalyanaraman, IEEE TPDS,
32(5):1191-1209, 2021.
P. Ghosh, S. Krishnamoorthy, A. Kalyanaraman, IPDPS'19, pp. 578-589, 2019.
Ripples(download)

Parallel software for influence maximization. Contains scalable multithreaded and MPI implementations for executing state-of-the-art approximation algorithms for the influence maximization problem.
Citations: M. Minutoli, M. Halappanavar, A. Kalyanaraman, A. Sathanur, R. Mcclure, J. McDermott. CLUSTER, p.12, 2019.
M. Minutoli, M. Dracco, M. Halappanavar, A. Tumeo, A. Kalyanaraman. ICS, 2020.
Hyppo-X(download)

An open source implementation for conducting topological data analysis on multi-dimensional point cloud data sets from phenomics and other domains. Tool supports automated and human-guided functions to visually navigate and extract topological interesting features toward hypotheses generation.
Citation: M. Kamruzzaman, A. Kalyanaraman, B. Krishnamoorthy, S. Hey, P. Schnable. TCBB'19, DOI: 10.1109/TCBB.2019.2947500.
biLouvain(download)
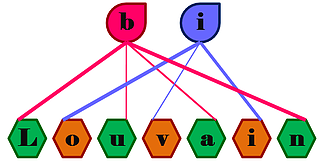
A C++ code for detecting communities in a bipartite network. There are versions of the code that can be used for strict bipartite community detection (using bipartite modularity as the objective), and for detecting communities in bipartite graphs that also have intra-type edges.
Citation: P. Pesantez, A. Kalyanaraman. IEEE TCBB, 2018, doi: 10.1109/TCBB.2017.2765319
A graph toolkit that contains multithreaded implementations for parallel graph community detection and for balanced graph coloring.
Citations: H. Lu et al. Parallel Computing, Vol.
47, pp. 19-37, 2015.
H. Lu et al. IEEE TPDS, Vol. 28, No. 5,
pp. 1240-1256, 2017.
pGraph-Tascel (download)
A parallel (MPI) program to construct large-scale protein sequence homology graphs on parallel distributed and shared memory computers. Given n input sequences, the goal is to identify all pairs of sequences that are highly similar (based on a user-specified set of alignment criteria). The code generates optimal alignments (using Smith-Waterman algorithm) and uses suffix trees to prune the search space prior to performing alignments. The implementation is parallel and runs on MPI clusters. It provides a scalable parallel implementation for all-against-all frameworks in bioinformatics.
Citation: J. Daily et al. JPDC, Vol. 79-80, pp. 132-142, 2015.
An older version of this method can be accessed here: download
Citation: C. Wu et al. IEEE TPDS, Vol. 23, No. 10, pp. 1923-1933, 2012.
pClust (download)
A scalable parallel software for detecting dense subgraphs
(clusters) in large-scale protein sequence homology graphs (e.g.,
output from pGraph).
Citation: C. Wu, A. Kalyanaraman, SC'08, pp. 1-10, 2008.
LTR_seq/LTR_par (download)
Serial and parallel software for de novo identification of full-length LTR retrotransposons (a class of genomic repeats).
Citation: A. Kalyanaraman, S. Aluru. JBCB, Vol. 4, No. 2, pp. 197-216, 2006.
PaCE (download)
Parallel software in C/MPI for clustering large-scale Expressed Sequence Tags (or ESTs) data. This tool was developed in the early 2000s. It can also be used for modern day RNAseq/transcriptomics data clustering although it is recommended that the new generation tools be used for that purpose.
Citation: A. Kalyanaraman et al., NAR, Vol. 31, No. 11, pp. 2963-2974, 2003.

